Clustering
K-means
Given a set of $n$ observations, k-means clustering aims to partition the $n$ observations into $k\leq n$ sets, to minimize the within-cluster sum of squares (i.e. variance). The objective is:
\[\text{argmin}_{S} \sum_{i=1}^k \sum_{x\in S_i} \| x - \mu_i\|^2 = \text{argmin}_{S} \sum_{i=1}^k |S_i| \text{Var}(S_i)\] \[\text{where} \quad \mu_i = \frac{1}{|S_i|} \sum_{x \in S_i}x \quad \small{\text{is the centroid of points in }} S_i\]Lloyd’s algorithm :
- Initialize: Pick K centroids $m_1, m_2, \dots, m_k$
- Repeat until convergence:
- Asssignment step: Assign each observation to the cluster with the nearest mean (centroid) \(S_i = \big\{ x_p: \| x_p - m_i\|^2 \leq \| x_p - m_j\|^2 \quad \forall j, 1\leq j\leq k \big\}\)
- Update step: Recalculate means (centroids) for observations assigned to each cluster \(m_i = \frac{1}{|S_i|} \sum_{x_j \in S_i} x_j\)
Lloyd’s algorithm has guaranteed finite convergence to local minimum, as the loss $J$ (see below) decreases monotonically, is bounded below by $0$ and the partition space is finite (at most $K^n$ possible assignments).
Mean is the Optimal Centroid
\(\textit{Proof:}\)
Let $J = \sum_{i=1}^k \sum_{x\in S_i} | x - \mu_i|^2$
Take derivative:
Set to 0:
\[\sum_{x\in S_i} (x_i - \mu_i) = 0 \iff \sum_{x\in S_i} x_i = |S_i|\mu_i \iff \mu_i = \frac{1}{|S_i|}\sum_{x\in S_i} x_i\]Initialization methods
- Random Partition: assign each point randomly to a cluster, compute means
-
Forgy Method: pick K random points as initial centroids
- K-means++: spreads out the initial cluster centers. Cluster centers are sampled sequentially and each new cluster is chosen from the remaining data points with probability proportional to its square distance from the closest existing cluster center.
- $m_1 \sim \mathrm{Uniform}({x_1, x_2, \dots, x_n})$
- for $t = 2, \dots, k$:
- for each point $x_i$:
- $d(x_i) = \min_{j < t} |x_i - m_j|^2$
- $P(x_i \text{ selected}) = \frac{d(x_i)}{\sum_j d(x_j)}$
- for each point $x_i$:
- Farthest Point: deterministic, always pick farthest point from current centroids
Variants
- K-medians: minimize $\sum_{i=1}^k \sum_{x\in S_i} | x - \mu_i|$. More robust to outliers
- K-medoids: has the constraint that $\mu_k$ must be an actual data point
- Fuzzy C-means: soft clustering approach, minimize $\sum_{i=1}^k \sum_{j} w_{ji}^m || x_j - \mu_i||^2$ where each element $w_{ji}$ tells the degree to which element $x_j$ belongs to cluster $S_i$. $w_{ji} = \Big[\sum_{i=1}^k\big(\frac{ | x_j - \mu_i|}{| x_j - \mu_k|} \big)^{\frac{2}{m-1}}\big]^{-1} $
- Kernel K-means: Operate in implicit feature space $\psi(x)$
- Spherical K-means: use cosine similarity instead of euclidean. Normalize updates $\mu_k = (\Sigma x_i) / | \Sigma x_i |$
Computational upgrades
- Vectorization: $|x_i - \mu_k|^2 = x_i^\top x_i - 2x_i^\top\mu_k + \mu_k^\top\mu_k$ which can be computed as matrix operations. So we can compute all pairwise squared distances efficiently and don’t need to loop over: observations, clusters, dimensions.
- Mini-Batch K-means: $\mu_k^{new} =\mu_k + \frac{b}{v_k + b} \big( \frac{1}{b} \sum_{i=1}^b x_i - \mu_k\big)$ if we’ve seen $v_k$ points in cluster $k$ so far and we assign $b$ new points to that cluster.
Problems with K-Means
- Cluster of similar expected size. This is the assumption of spherical cluster that are separable so that the mean converges towards the geometric cluster center. So K-means works poorly on elongated and concentric clusters and clusters of different density.
- Can only find convex clusters.
- Sensitive to initialization. Different starting points can yield very different final clusters. So can converge to a local minimum.
- Sensitive to scale. Requires careful normalization/standardization otherwise features with large variance dominate the distance calculation.
- Assumes equal importance to each feature.
- Sensitive to outliers and noise.
- Computationally expensive: $O(n\times k\times d \times iterations)$.
- Curse of dimensionality: all pairwise distance becomes similar (concentration phenomenon).
Hierarchical Clustering
Hierarchical clustering provides an alternative view on K–means in the sense that it does not require us to pre-specify the number of clusters $K$. It results in a tree-based representation of the observations, called a dendogram. It also requires to specify a measure of (dis)similarity between groups of observations. Hierarchical clustering can be done in two manners: \texit{agglomerative} or divisive. Agglomerative recursively merge a selected pair of clusters into a single one, where the pair chosen consists of the two groups with the smallest pairwise dissimilarity. Divisive methods start with the full dataset and recursively split one cluster into two new.
The extent to which the hierarchical structure produced by a dendogram actually fits the data can be judged by the cophenetic correlation coefficient, which is the correlation between all the $N(N-1)/2$ pairwise dissimilarities $d(i,j)$ and their cophenetic dissimilarities $C(i,j)$ produced by the dendogram.
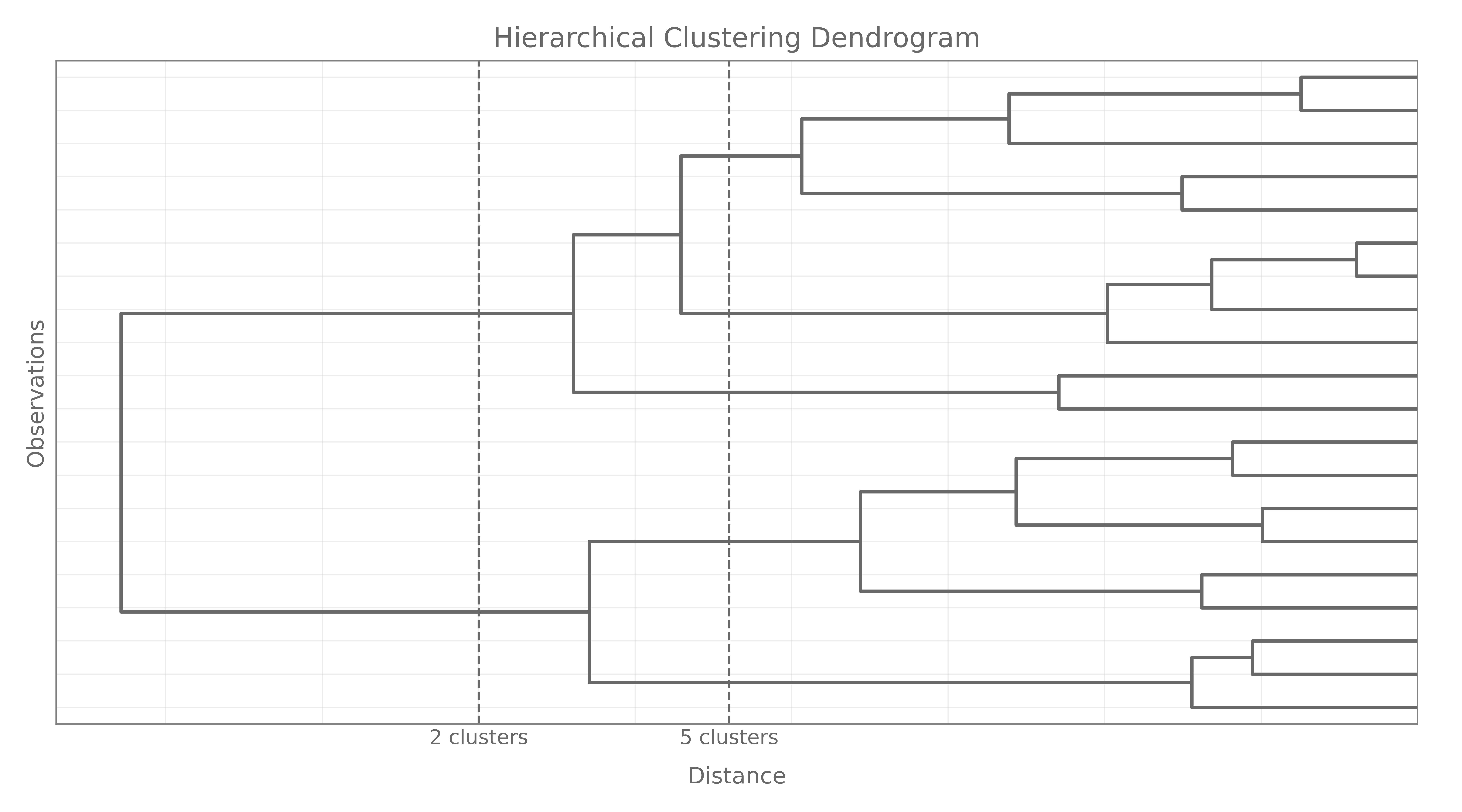
Cluster Linkage
- Maximum / Complete linkage: \(\max_{a\in A, b \in B} d(a,b)\)
- Minimum / single linkage (nearest neighbour): \(\min_{a\in A, b \in B} d(a,b)\)
- Average linkage: $\frac{1}{ | A | \cdot | B |} = \sum_A \sum_B d(a,b)$
- Centroid linkage: \(d(\mu_A, \mu_B)\)
- Ward’s Method (Minimum Variance): $\frac{|A|\cdot |B|}{|A|+|B|} || \mu_a - \mu_b||^2$
Choice of Distance functions
- Minkowski Distance: $d(x,y) = \Big( \sum_i | x_i - _i|^q\Big)^{1/q} $
- Euclidean Distance for $q = 2$
- Manhattan Distance for $q = 1$
- Chebyshev Distance for $q \to \infty$
- Mahalanobis Distance: \(d(x,y) = \sqrt{(x-y)^\top \Sigma^{-1}(x-y)}\)
- Covariance aware: computes ordinary Euclidean distance in the correlation and variance corrected space
- Requires estimation of covariance matrix so can be unstable in high dimensions, and expensive
- Cosine Distance ($1 -$ Cosine Similarity): \(d(x,y) = 1 - \frac{x \cdot y}{\|x\| \|y\|}\)
- Excellent for high dimensional sparse vector
- Hamming distance (for Categorical Data): \(d(x,y) = \sum_i I(x_i \neq y_i)\)
- Dynamic Time Warping (for Time Series):
- First build a cost matrix where $d$ is usually the Euclidean distance: $C(t_1, t_2) = d(X_{t_1}, Y_{t_2})$
- Calculate cumulative path with Dynamic Programming: \(D(t_1, t_2) = C(t_1, t_2) + \min \Big\{ D(t_1-1, t_2), D(t_1, t_2-1), D(t_1-1, t_2-1)\Big\}\)
-
the DTW distance is the value for the final time indices. The path taken by the dynamic programming table is called the warping path
- Works even when time series are misaligned, different speeds or unequal lenghts
- Very effective for clustering and classification
- Computation is $O(n \times m)$ where $n$ and $m$ are the length of each series, so can be slow.
- Not a true metric (as triangle inequality fails)
Metrics and Number of Clusters
Clustering Metrics
- WCSS / Inertia (Within-Cluster Sum of Squares): measures the total square distance from each point to its cluster center: \(WCSS = \sum_{i=1}^k \sum_{x\in S_i} \| x - \mu_i\|^2\) use it to compare different methods with the same number of clusters.
- BCSS (Between-Cluster Sum of Squares) \(BCSS = \sum_{i\neq j}^k \|\mu_i - \mu_j \|^2\)
- TSS (Total Sum of Squares) $TSS = WCSS + BCSS$
- Silhouette Coefficient (Most popular internal metric): For each point $i$:
- $a(i) = $ average distance from i to all other points in the same cluster
- $b(i) = $ minimum average distance from i to points in any other cluster
- $s(i) \approx 1$ tells that the point is well clustered (far from other points), $s(i) \approx 0$ that the point is on the boundary between clusters, and $s(i) \approx -1$ that the point is probably in the wrong cluster
- $Overall \hspace{0.5em} Silhouette \hspace{0.5em} Score = \frac{1}{n} \sum s(i)$
- Works with any distance metrics. However it has $O(n^2)$ computation, is biased towards complex clusters and can be sensitive to noise and outliers.
- Silhouette plot generally shows the histogram of silhouettes per cluster
- Davies-Bouldin Index:
\(DB = \frac{1}{n} \sum_{i=1}^k \max_{i \neq j} \bigg( \frac{\sigma_i + \sigma_j}{d(\mu_i, \mu_j)}\bigg)\)
- $\sigma_i$ is the average distance of all elements in cluster $i$ to its centroid $\mu_i$
- $d(\mu_i, \mu_j)$ is the distance between centroids
- The range is $[0, \infty)$ and $0$ is optimal
- $\max_{i \neq j} \big( \frac{\sigma_i + \sigma_j}{d(\mu_i, \mu_j)}\big)$ is the worst-case similarity of cluster $i$ to another cluster, and $DB$ is the average worst-case similarity across all clusters
- $O(nK)$, but biased towards spherical clusters, sensitive to outliers, and might not work well with different density clusters
- Dunn Index:
\(D = \frac{\min_{1\leq i < j \leq k} d(i,j)}{\max_{1 \leq c \leq k} d'(c)}\)
- $d(i,j)$ is the distance between clusters $i$ and $j$, which can be any number of distance measure, such as the distance between the centroids of the clusters
- $d’(c)$ is a measure of intra-cluster distance for cluster $c$. It can also be measured any metric, like the maximal distance between any two elements in cluster $c$
- The range is $[0, \infty)$ and higher is better
- $O(n^2)$ Complexity, extremely sensitive to outliers (numerator and denominator) and noise
Choosing the number of Clusters
- Elbow Method:
- The Elbow method consists of choosing a range of $k$ values, and computing the $WCSS$ for each $k$, and plotting it. Look for elbow where the rate of decrease sharply drops
- Works as $WCSS$ always decreases with higher $k$
- However can have some problems, like no clear elbow or multiple elbows.
- One can use the Kneedle Algorithm, which is an automatic detection algorithm, or the Second Derivative method
- Silhouette Analysis: Can theoretically work with other metrics but Silhouette score is always the preferred one. Just plot Silhouette vs. number of clusters and pick the highest one.
- Dendogram: Usually done for Hierarchical Clustering
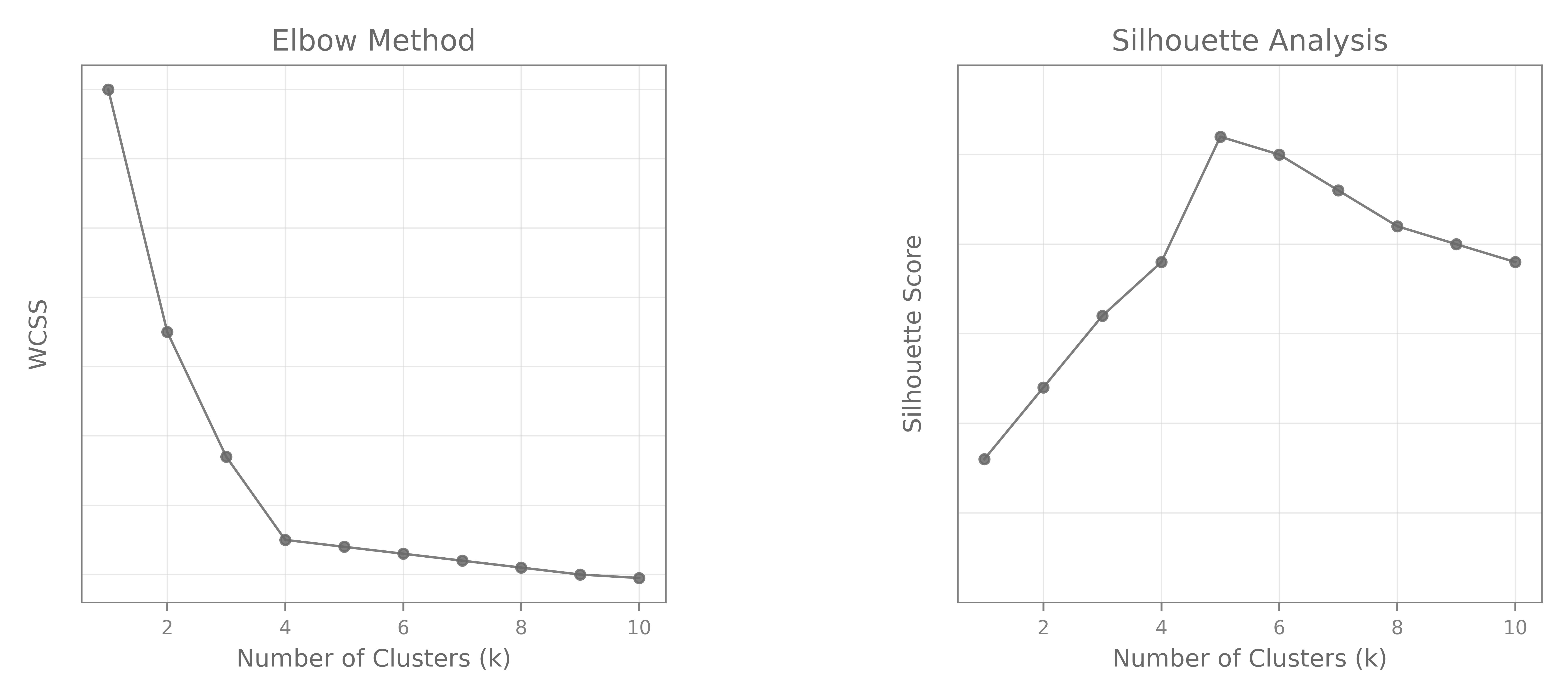
Note: This is not supposed to represent plots for the same dataset / clustering method. Those are just illustrative. And yes, the right choice looks like $K=4$ for the Elbow plot, and $K=5$ for the Silhouette
Gaussian Mixture Models
GMM is a probabilistic model assuming the data is generated from a mixture of $K$ Gaussian distribution with unkownd parameters. The model takes the following form: \(p(x) = \sum_{k=1}^K \pi_k \cdot \mathcal{N}(x |\mu_k, \Sigma_k)\)
- Soft clustering: each point has a (posterior) probability of belonging to each cluster
- Flexible cluster shapes (can have different orientations, sizes)
- Assumes Gaussian (fails it data isn’t $\approx \mathcal{N})$
- Sensitive so initialization
- Requires to specify $K$ upfront
- Expensive in parameters and has requires a lot of data as can be unstable in high dimensions
Likelihood to maximize:
\[L(\theta) = \prod_{i=1}^n \sum_{k=1}^K \pi_k\, \mathcal{N}(x_i \mid \mu_k, \Sigma_k) \qquad \iff \qquad \ell(\theta) = \sum_{i=1}^n \log\!\left( \sum_{k=1}^K \pi_k\, \mathcal{N}(x_i \mid \mu_k, \Sigma_k) \right)\]The presence of the logarithm of a sum prevents closed-form parameter updates, making direct maximization intractable.
Posterior Probabilities (Soft Assignments):
Using Bayes’ rule, the responsibility of component $k$ for point $x_i$ is:
Complete-Data Log-Likelihood:
If the latent assignment indicators $z_{ik}$ were known:
The EM algorithm optimizes the expected complete-data log-likelihood:
Expectation Maximization
The EM algo is an 2-step iterative optimization technique to estimat ethe unkown parameters in probabilistic models.
-
E-Step (Expectation Step):
\[\gamma_{ik}^{(t)} = P(z_i = k \mid x_i, \theta^{(t)}) = \frac{\pi_k^{(t)}\, \mathcal{N}(x_i \mid \mu_k^{(t)}, \Sigma_k^{(t)})} {\sum_{j=1}^K \pi_j^{(t)}\, \mathcal{N}(x_i \mid \mu_j^{(t)}, \Sigma_j^{(t)})}.\]- Compute the posterior “responsibility” that component $k$ generated point $x_i$ by using the current parameter estimates $\theta^{(t)} = {\pi_k^{(t)},\mu_k^{(t)},\Sigma_k^{(t)}}$.
- Converts latent assignments $z_{ik}$ into soft probabilities $\gamma_{ik}$, i.e. expected values of the hidden variables $z_{ik}$.
-
M-Step (Maximization Step)
\[\pi_k^{(t+1)} = \frac{1}{n}\sum_{i=1}^n \gamma_{ik}^{(t)}, \qquad \mu_k^{(t+1)} = \frac{\sum_{i=1}^n \gamma_{ik}^{(t)} x_i}{\sum_{i=1}^n \gamma_{ik}^{(t)}},\] \[\Sigma_k^{(t+1)} = \frac{\sum_{i=1}^n \gamma_{ik}^{(t)} (x_i - \mu_k^{(t+1)})(x_i - \mu_k^{(t+1)})^\top} {\sum_{i=1}^n \gamma_{ik}^{(t)}}\]- Update mixture weights, means, and covariance matrices using $\gamma_{ik}^{(t)}$, assuming those the soft assignments are correct. These are basically the Maximum Likelihood Estimates of the parameters, given the soft-assignments.
- Maximizes the expected complete-data log-likelihood: $ Q(\theta \mid \theta^{(t)}) = \mathbb{E}_{z|X,\theta^{(t)}}[\ell_c(\theta)]. $
When I was an undergraduate student, I always found the Expectation-Maximization algorithm very unintuitive. I learned it many times, but it never stuck, because I never truly understood it. I’m still no pro of that EM procedure, but maybe this will help an unfortunate student that wished his exam was on supervised learning only.
It turns out that LLoyd’s Algorithm for K-means is a special case of the Expectation Maximization procedure. Actually, under certain (very strict) conditions, the two are equivalent. You would essentially need each component to have equal, spherical covariance, the variances going to 0, each cluster having equal mixing weight (i.e. equal $\pi_k$), and the procedure itself to be a hard assignment method instead of a soft, probabilistic algorithm. This doesn’t help much does it?
What if I told you the E-Step was equivalent to the Assignment Step and the M-Step to the Update Step? One could see that the E-step calculates the expected membership probabilities for (or associates the cluster center to) each data point under the current parameter estimates (or centroids positions), and the M-Step updates the model parameters (centroid positions) given the current membership probabilities (cluster assignments).
Convergence
- The observed (expected) log-likelihood increases monotonically (actually non-decreasing).
- EM converges to a stationary point, however it might be a saddle point or a local maximum
This is all for today. Actually, this is way too much for any day.
References
The Elements ofStatistical Learning: Data Mining, Inference, and Prediction. (2009)
Trevor Hastie, Robert Tibshirani, Jerome Friedman
Book
k-means++: The Advantages of Careful Seeding (2006)
David Arthur, Sergei Vassilvitskii
Paper
Maximum Likelihood from Incomplete Data via the EM Algorithm (1977)
A. P. Dempster, N. M. Laird, D. B. Rubin
Paper
The EM Algorithm and Extensions (2008)
GJ McLachlan, T Krishnan
Book