Dependency and Discrepancy Measures in RKHS
This blog post provides an introduction to kernel methods in modern statistical learning, starting from the foundational theory of reproducing kernel Hilbert spaces (RKHS), then developing on the theory of mean embeddings and covariance operators. Three major kernel-based hypothesis testing frameworks: Maximum Mean Discrepancy, the Hilbert-Schmidt Independence Criterion, and the Kernel Stein Discrepancy are introduced.
1. Primer on Kernels
1.1 RKHS and Properties of Kernels
This section aims to be an introduction to kernel functions, reproducing kernel Hilbert spaces (RKHS), and their main theorems and properties, which are direct results of functional analysis. It is intended to be introductory and to present these concepts from a theoretical standpoint. Subsequent sections will emphasize applied and practical aspects of kernel methods in statistical frameworks. For a more complete and rigorous treatment, we refer to Smola & Schölkopf (1998) and Berlinet & Thomas-Agnan (2011).
1.1.1 Kernel
Definition (Kernel). A function $k : \mathcal{X} \times \mathcal{X} \rightarrow \mathbb{R}$ is a kernel if it is symmetric and positive semi-definite: for any $n \in \mathbb{N}$, points ${x_1, \ldots, x_n} \subset \mathcal{X}$, and coefficients $c_1, \ldots, c_n \in \mathbb{R}$,
\[\sum_{i=1}^{n} \sum_{j=1}^{n} c_i c_j k(x_i, x_j) \geq 0.\]Equivalently, the Gram matrix $\mathbf{K} \in \mathbb{R}^{n \times n}$ with $K_{ij} = k(x_i, x_j)$ is positive semi-definite for any finite point set.
Remark. Every kernel admits a feature map representation $k(x, x’) = \langle \phi(x), \phi(x’) \rangle_{\mathcal{H}}$ for some Hilbert space $\mathcal{H}$, which will be formalized via the associated RKHS.
Example. Standard kernels on $\mathbb{R}^d$ include the linear kernel $k(x, x’) = x^\top x’$, the polynomial kernel $k(x, x’) = (x^\top x’ + c)^p$, the RBF kernel $k(x, x’) = \exp(-|x - x’|^2/2\sigma^2)$, and the Laplacian kernel $k(x, x’) = \exp(-|x - x’|/\sigma)$.
Proposition (Closure and composition properties). Kernels are closed under positive linear combinations ($\alpha k_1 + \beta k_2$ for $\alpha, \beta \geq 0$), products ($k_1 \cdot k_2$), pointwise limits, and tensor products. Moreover, if $k$ is a kernel on $\mathcal{X}$ and $\psi: \mathcal{Y} \rightarrow \mathcal{X}$, then $k(\psi(\cdot), \psi(\cdot))$ is a kernel on $\mathcal{Y}$. Products preserve universality and characteristic properties.
1.1.2 RKHS
Definition (Hilbert Space). A Hilbert space $\mathcal{H}$ is a complete inner product space, i.e., a vector space equipped with an inner product $\langle \cdot, \cdot \rangle_{\mathcal{H}}$ such that every Cauchy sequence converges in the induced norm \(\|f\|_{\mathcal{H}} = \sqrt{\langle f, f \rangle_{\mathcal{H}}}\).
Definition (Reproducing Kernel Hilbert Space). A Hilbert space $\mathcal{H}$ of real-valued functions $f: \mathcal{X} \rightarrow \mathbb{R}$ is a reproducing kernel Hilbert space (RKHS) if point evaluation functionals are continuous. That is, for each $x \in \mathcal{X}$, the evaluation functional $\delta_x: \mathcal{H} \rightarrow \mathbb{R}$ defined by $\delta_x(f) = f(x)$ is bounded.
Theorem (Riesz Representation). Let $\mathcal{H}$ be a Hilbert space and $L: \mathcal{H} \rightarrow \mathbb{R}$ a bounded linear operator. Then there exists a unique element $h \in \mathcal{H}$ such that $Lf = \langle f, h \rangle_{\mathcal{H}}$ for all $f \in \mathcal{H}$.
Proposition (Reproducing kernel). By the Riesz representation theorem, for each $x \in \mathcal{X}$, there exists a unique element $k(\cdot, x) \in \mathcal{H}$ such that
\[f(x) = \langle f, k(\cdot, x) \rangle_{\mathcal{H}} \quad \text{for all } f \in \mathcal{H}.\]This is the reproducing property. The function $k: \mathcal{X} \times \mathcal{X} \rightarrow \mathbb{R}$ defined by $k(x, x’) = \langle k(\cdot, x), k(\cdot, x’) \rangle_{\mathcal{H}}$ is called the reproducing kernel of $\mathcal{H}$. The kernel $k$ is uniquely determined by $\mathcal{H}$, and conversely, $\mathcal{H}$ is uniquely determined by $k$.
Theorem (Moore-Aronszajn). There is a one-to-one correspondence between kernels on $\mathcal{X}$ and RKHSs of real-valued functions on $\mathcal{X}$. Given any kernel $k$, there exists a unique RKHS $\mathcal{H}_k$ with reproducing kernel $k$. Conversely, every RKHS uniquely determines its reproducing kernel.
Remark (The RKHS norm). The RKHS norm can be interpreted through the reproducing property. For \(f \in \mathcal{H}_k\), we can write $f$ as a (possibly infinite) linear combination of kernel evaluations. The norm $|f|_{\mathcal{H}_k}$ measures the complexity or smoothness of $f$ in a way that depends on the choice of kernel $k$. Functions that align with the kernel structure have smaller norms.
Definition (Kernel integral operator). Let $\mu$ be a measure on $\mathcal{X}$. The kernel integral operator $T_k: L^2(\mathcal{X}, \mu) \rightarrow L^2(\mathcal{X}, \mu)$ associated with kernel $k$ is defined by
\[(T_k f)(x) = \int_{\mathcal{X}} k(x, x') f(x') \, d\mu(x').\]Under appropriate conditions, $T_k$ is a compact, self-adjoint, positive operator.
Theorem (Mercer). Let $\mathcal{X} \subset \mathbb{R}^d$ be compact, $\mu$ a finite Borel measure with support $\mathcal{X}$, and $k: \mathcal{X} \times \mathcal{X} \to \mathbb{R}$ a continuous positive definite kernel. Then the integral operator $T_k$ on $L^2(\mathcal{X}, \mu)$ is compact, self-adjoint, and positive, with a countable set of eigenvalues \(\{\lambda_i\}_{i=1}^{\infty}\) (with $\lambda_i \geq 0$, $\lambda_i \to 0$) and corresponding orthonormal eigenfunctions ${\phi_i}_{i=1}^{\infty}$ forming an orthonormal basis of $L^2(\mathcal{X}, \mu)$. The kernel admits the absolutely and uniformly convergent representation
\[k(x, x') = \sum_{i=1}^{\infty} \lambda_i \phi_i(x) \phi_i(x'), \quad \forall x, x' \in \mathcal{X}.\]The RKHS \(\mathcal{H}_k\) consists of all functions $f \in L^2(\mathcal{X}, \mu)$ with expansion $f = \sum_{i=1}^{\infty} \alpha_i \phi_i$ satisfying
\[\|f\|_{\mathcal{H}_k}^2 = \sum_{i=1}^{\infty} \frac{\alpha_i^2}{\lambda_i} < \infty,\]where $\alpha_i = \langle f, \phi_i \rangle_{L^2(\mathcal{X}, \mu)}$. This norm follows from the orthogonal expansion and the identity \(\langle \phi_i, \phi_j \rangle_{\mathcal{H}_k} = \delta_{ij}/\lambda_i\).
Note that any function in the RKHS $\mathcal{H}$ can be written as a linear combination of the eigenfunctions, via Mercer’s theorem, i.e. $f = \sum_{i=1}^{\infty} \alpha_i \phi_i(x) \quad \forall f \in \mathcal{H}_k$.
Remark (Isometric views of the RKHS). Mercer’s theorem reveals three equivalent perspectives on the RKHS:
- The function space view: $\mathcal{H}_k$ as a space of functions with the reproducing property
- The feature space view: Functions as elements in a potentially infinite-dimensional Hilbert space with feature map $\phi: x \mapsto (\sqrt{\lambda_i}\phi_i(x))_{i=1}^{\infty}$, where $k(x, x’) = \langle \phi(x), \phi(x’) \rangle$
- The sequence space view: $\mathcal{H}_k$ is isometrically isomorphic to the weighted $\ell^2$ space $\mathcal{H}_k \cong \ell^2(\lambda) = {(\alpha_i): \sum_i \alpha_i^2/\lambda_i < \infty}$
Remark (The kernel trick). In practice, kernels provide a powerful analytical tool for performing comparisons through inner products in potentially infinite-dimensional feature spaces, without ever having to explicitly augment the data or compute the feature map $\phi$. Given data points $x_1, \ldots, x_n \in \mathcal{X}$, we can compute inner products in the feature space simply by evaluating $k(x_i, x_j)$. This “kernel trick” allows algorithms that depend only on inner products (such as ridge regression, SVMs, and PCA) to be kernelized, operating implicitly in high- or infinite-dimensional spaces while maintaining computational tractability. The choice of kernel thus encodes prior knowledge about what constitutes similarity in the problem domain.
Proposition (Derivative reproducing). Let $k$ be a smooth kernel such that all partial derivatives $\partial^{\alpha} k(x, x’)$ exist and are continuous. If $f \in \mathcal{H}_k$ is sufficiently smooth, then the derivatives of $f$ can be computed via
\[\partial^{\alpha} f(x) = \langle f, \partial^{\alpha}_x k(\cdot, x) \rangle_{\mathcal{H}_k},\]where the derivative is taken with respect to the first argument of $k$. This extends the reproducing property to derivatives, which is particularly useful in operator learning and solving differential equations in RKHS.
1.2 Mean Embedding
Definition (Kernel Mean Embedding). Given a kernel $k$ with reproducing kernel Hilbert space (RKHS) \(\mathcal{H}_k\), the kernel mean embedding of a probability measure \(\mathbb{P} \in \mathcal{M}^1_+(\mathcal{X})\) is the element \(\mu_\mathbb{P} \in \mathcal{H}_k\) defined by
\[\mu_\mathbb{P} = \mathbb{E}_{X \sim \mathbb{P}}[k(X, \cdot)] = \int_{\mathcal{X}} k(x, \cdot) \, \mathrm{d}\mathbb{P}(x)\]Using the reproducing property of \(\mathcal{H}_k\), \(\mu_\mathbb{P}\) is the unique element of \(\mathcal{H}_k\) satisfying \(\langle f, \mu_\mathbb{P} \rangle_{\mathcal{H}_k} = \mathbb{E}_{X \sim \mathbb{P}}[f(X)] \quad \forall f \in \mathcal{H}_k\).
In particular, for $f = k(\cdot, x)$, this yields \(\mu_\mathbb{P}(x) = \langle \mu_\mathbb{P}, k(\cdot, x) \rangle_{\mathcal{H}_k} = \mathbb{E}_{X \sim \mathbb{P}}[k(X, x)]\).
Given an i.i.d. sample ${x_1,\dots,x_m}$ from $\mathbb{P}$, the empirical kernel mean embedding is
\[\hat{\mu}_\mathbb{P} = \frac{1}{m} \sum_{i=1}^m k(x_i, \cdot)\]
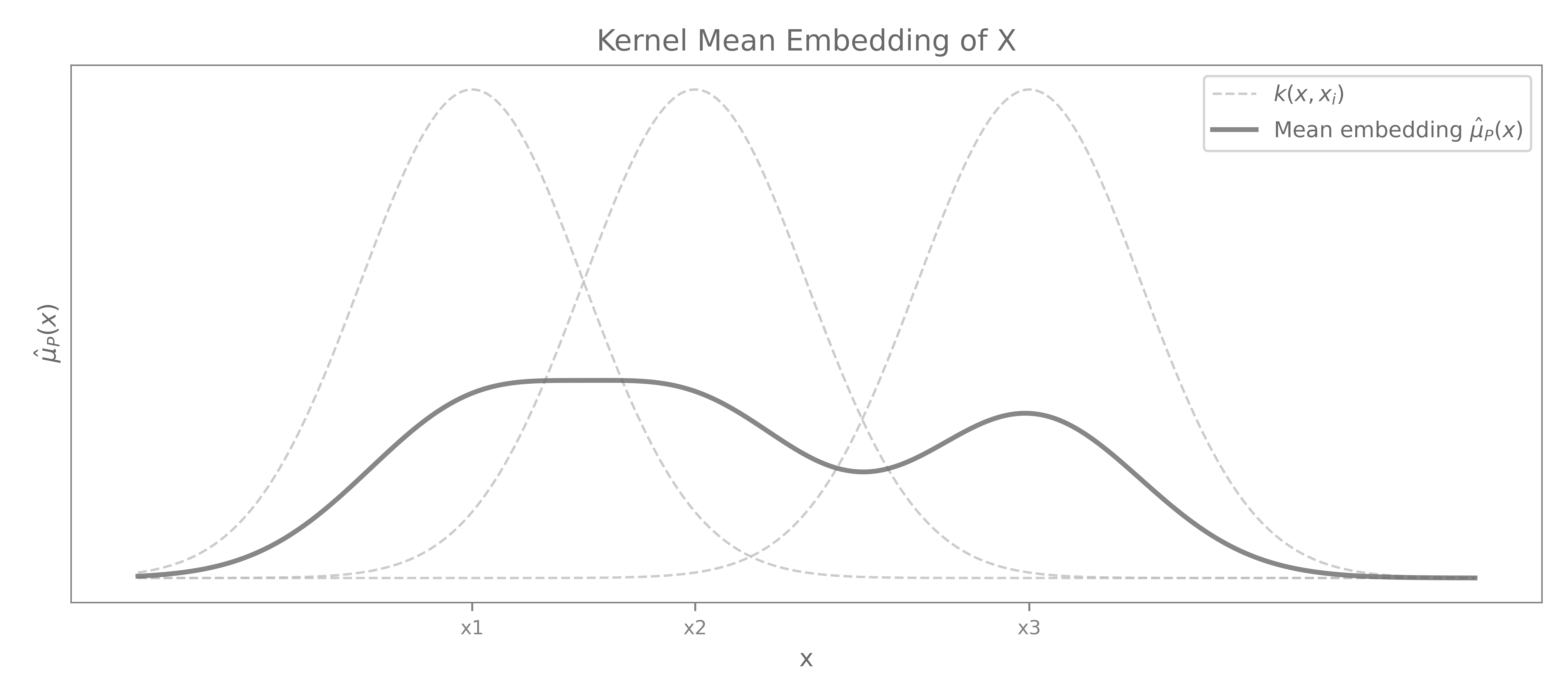
Remark (Existence). The embedding \(\mu_\mathbb{P}\) exists in \(\mathcal{H}_k\) when \(\mathbb{E}_{X \sim \mathbb{P}}[\sqrt{k(X,X)}] < \infty\), which holds automatically for bounded kernels.
Definition (Characteristic Kernel). A kernel $k$ is characteristic if the mean embedding map $\mathbb{P} \mapsto \mu_\mathbb{P}$ is injective, meaning $\mu_\mathbb{P} = \mu_\mathbb{Q}$ if and only if $\mathbb{P} = \mathbb{Q}$.
1.3 Tensor Product Space and Covariance Operators
1.3.1 Tensor Product Space
Let $(\mathcal{H}, k)$ and $(\mathcal{G}, \ell)$ be two reproducing kernel Hilbert spaces over input spaces $\mathcal{X}$ and $\mathcal{Y}$ respectively. The tensor product space $\mathcal{G} \otimes \mathcal{H}$ provides a natural framework for studying joint distributions and dependence structures between random variables taking values in $\mathcal{X}$ and $\mathcal{Y}$.
Definition (Tensor Product RKHS). The tensor product space $\mathcal{G} \otimes \mathcal{H}$ is the completion of the linear span of elementary tensors ${g \otimes h : g \in \mathcal{G}, h \in \mathcal{H}}$ under the inner product
\[\langle g_1 \otimes h_1, g_2 \otimes h_2 \rangle_{\mathcal{G} \otimes \mathcal{H}} = \langle g_1, g_2 \rangle_{\mathcal{G}} \langle h_1, h_2 \rangle_{\mathcal{H}}.\]The space $\mathcal{G} \otimes \mathcal{H}$ is itself an RKHS over $\mathcal{Y} \times \mathcal{X}$ with reproducing kernel $k \otimes \ell$:
\[(k \otimes \ell)((y,x), (y',x')) = k(x,x') \ell(y,y').\]Any functional tensor $g \otimes h$ can be interpreted as a linear operator from $\mathcal{H}$ to $\mathcal{G}$ via the mapping
\[(g \otimes h)(f) = \langle h, f \rangle_{\mathcal{H}} g \quad \text{for } f \in \mathcal{H}.\]This operator takes an element $f \in \mathcal{H}$, computes its inner product with $h$ in the RKHS $\mathcal{H}$ (yielding a scalar), and scales the element $g \in \mathcal{G}$ by this scalar. Using the reproducing property with feature maps $\phi(y) = \ell(y, \cdot) \in \mathcal{G}$ and $\Phi(x) = k(x, \cdot) \in \mathcal{H}$, we have
\[(\phi(y) \otimes \Phi(x))(f) = \langle \Phi(x), f \rangle_{\mathcal{H}} \phi(y) = f(x) \phi(y).\]For any $g, f \in \mathcal{G}$ and $u, v \in \mathcal{H}$,
\[\langle g, (u \otimes v)(f) \rangle_{\mathcal{G}} = \langle g, \langle v, f \rangle_{\mathcal{H}} u \rangle_{\mathcal{G}} = \langle v, f \rangle_{\mathcal{H}} \langle g, u \rangle_{\mathcal{G}} = \langle u \otimes v, g \otimes f \rangle_{\mathcal{G} \otimes \mathcal{H}}.\]This identity shows that evaluating the operator $u \otimes v$ on $f$ and taking the inner product with $g$ in $\mathcal{G}$ is equivalent to computing the tensor product inner product of $u \otimes v$ with $g \otimes f$ in $\mathcal{G} \otimes \mathcal{H}$.
Definition (Hilbert-Schmidt Operators). Let the bounded linear operator \(C: \mathcal{H} \to \mathcal{G}\) on separable Hilbert Spaces \(\mathcal{H}\) and \(\mathcal{G}\) with orthonormal basis \((h_i)_{i \in I}\) and \((g_j)_{j \in J}\) respectively. Define the Hilbert–Schmidt norm of $C$ by
\[\begin{align*} \|C\|_{\mathrm{HS}}^{2} &= \sum_{i \in I} \|C h_i\|_{\mathcal{G}}^{2} \\ &= \sum_{i \in I} \sum_{j \in J} \big| \langle C h_i , g_j \rangle_{\mathcal{G}} \big|^{2}. \end{align*}\]The Hilbert-Schmidt norm \(\|\cdot\|_{\text{HS}}\) is independent of the choice of orthonormal basis \(I, J\), which are assumed to be either finite or countably infinite. An operator $C$ is called a Hilbert–Schmidt operator if \(\|C\|_{\mathrm{HS}} < \infty\).
1.3.2 Covariance Operators
We now introduce the concepts of Cross-Covariance operator on reproducing kernel Hilbert spaces, as well as their normalized and conditional versions. The Cross-Covariance operator is a generalization of the covariance matrix to infinite dimensional feature spaces, used to capture dependence (and conditional dependence) of random variables under non-linear feature maps. The theoretical foundations and practical applications of Cross-Covariance operators have been introduced and thoroughly studied by K. Fukumizu, and we refer to his work for further details: Fukumizu et al. (2004, 2007, 2009). Most of the following definitions can be viewed as equivalent formulations of covariance, correlations and conditional correlations measures in Euclidean spaces generalized through non-linear kernels in reproducing kernel Hilbert spaces.
Let \(\mathcal{H}_\mathcal{X}\) and \(\mathcal{H}_\mathcal{Y}\) be two RKHS of functions on Borel sets $\mathcal{X}$ and $\mathcal{Y}$, with reproducing kernels $k_\mathcal{X}$ and $k_\mathcal{Y}$, respectively. Assuming integrability, i.e.
\[\mathbb{E}[k_\mathcal{X}(X,X)]< \infty \quad\forall X \in \mathcal{X}, \quad \text{and} \quad \mathbb{E}[k_\mathcal{Y}(Y,Y)]< \infty \quad\forall Y \in \mathcal{Y}\]we can define the cross-covariance operator of $(X,Y)$:
Definition (Cross-Covariance Operator). The centred cross-covariance operator of $(X,Y)$, denoted as $\Sigma_{YX}$, is an operator from \(\mathcal{H}_\mathcal{X}\) to \(\mathcal{H}_\mathcal{Y}\) defined, for all \(f \in \mathcal{H}_\mathcal{X}\) and \(g \in \mathcal{H}_\mathcal{Y}\), by:
\[\begin{align*} \langle g, \Sigma_{YX}f\rangle_{\mathcal{H}_\mathcal{Y}} &= \text{Cov}[f(X), g(Y)] \\ &= \mathbb{E}_{YX}[(f(X) - \mathbb{E}_{X \sim \mathbb{P}_X}[f(X)]) \cdot (g(Y) - \mathbb{E}_{Y \sim \mathbb{P}_Y}[g(Y)])] \\ &= \mathbb{E}_{YX}[\langle f, k_\mathcal{X}(X, \cdot) - \mu_{\mathbb{P}_X}\rangle \langle g, k_\mathcal{Y}(Y, \cdot) - \mu_{\mathbb{P}_Y}\rangle ] \\ &= \mathbb{E}_{YX}[\langle f \otimes g, (k_\mathcal{X}(X, \cdot) - \mu_{\mathbb{P}_X}) \otimes (k_\mathcal{Y}(Y, \cdot) - \mu_{\mathbb{P}_Y})\rangle ] \end{align*}\]Using the Riesz Representation Theorem, one can also write:
\[\Sigma_{YX} = \mathbb{E}_{YX}[(k_\mathcal{X}(X, \cdot) - \mu_{\mathbb{P}_X}) \otimes (k_\mathcal{Y}(Y, \cdot) - \mu_{\mathbb{P}_Y})]\]The assumptions of integrability implies that the cross-covariance operator is Hilbert Schmidt. The operator represents higher-order correlations of $X$ and $Y$ through feature maps $f(X)$ and $g(Y)$. Analogously to the variance in Euclidean spaces, if $X = Y$, $\Sigma_{XX}$ is called the covariance operator. The cross-covariance operator can be used to determine the independence of $X$ and $Y$: $\Sigma_{YX} = 0 \iff X$ and $Y$ are independent.
Definition (Normalized Cross-Covariance Operator). The normalized cross-covariance operator of $(X,Y)$, denoted as $V_{YX}$, is a unique bounded operator such that:
\[\Sigma_{YX} = \Sigma_{YY}^{1/2}V_{YX}\Sigma_{XX}^{1/2}\]The normalized cross-covariance operator can be seen as an analogue to the correlation, and similarly, its Hilbert Schmidt norm is less than or equal than 1.
Finally, if we consider another random variable $Z$ on \(\mathcal{Z}\), and the RKHS \(\mathcal{H}_{\mathcal{Z}}\) with corresponding kernel \(k_\mathcal{Z}\), we can define the normalized conditional cross-covariance operator.
Definition (Normalized Conditional Cross-Covariance Operator). The normalized conditional cross-covariance operator of $X$ and $Y$ given $Z$, denoted as \(V_{YX\|Z}\), is defined as:
\[V_{YX|Z} = V_{YX} - V_{YZ}V_{XZ}\]Alternatively, it can also be defined as:
\[V_{YX|Z} = \Sigma_{YY}^{-1/2} \Sigma_{YX|Z} \Sigma_{XX}^{-1/2} \qquad \text{with} \qquad \Sigma_{YX|Z} = (\Sigma_{YX} -\Sigma_{YZ}\Sigma_{ZZ}^{-1}\Sigma_{ZX})\]Similarly to the cross-covariance operator, we have \(\Sigma_{YX\|Z}= 0 \iff X\) and $Y$ are independent given $Z$.
Empirically, given i.i.d samples \((x_i, y_i)_{i=1}^m\), we can denote \(k_\mathcal{X}(x_i, \cdot) = \varphi(x_i)\) and \(k_\mathcal{Y}(y_i, \cdot) = \psi(y_i)\). Now define
\[\hat{\mu}_X = \frac{1}{m} \sum_{i=1}^m \varphi(x_i) \qquad \text{and} \qquad \hat{\mu}_Y = \frac{1}{m} \sum_{i=1}^m \psi(y_i)\]the empirical kernel mean embeddings of $X$ and $Y$. Then the empirical cross-covariance operator is
\[\widehat{\Sigma}_{YX} = \frac{1}{m} \sum_{i=1}^m (\varphi(x_i) - \hat{\mu}_X) \otimes (\psi(y_i) - \hat{\mu}_Y)\]We often see the empirical cross-covariance operator denoted in matrix form, using \(\Phi = [\varphi(x_1), \dots, \varphi(x_m)]\) and \(\Psi = [\psi(y_1), \dots, \psi(y_m)]\) the feature matrices in \(\mathcal{H}_{\mathcal{X}}\) and \(\mathcal{H}_{\mathcal{Y}}\) respectively, and \(H = I - \frac{1}{m}\mathbf{1}\mathbf{1}^\top\) the centering matrix. Then
\[\widehat{\Sigma}_{YX} = \frac{1}{m} \Phi H \Psi^\top\]2. Testing with Kernels
2.1 Maximum Mean Discrepancy (MMD)
2.1.1 Theory
The Maximum Mean Discrepancy is a statistic to test whether two distributions $\mathbb{P}$ and $\mathbb{Q}$ differ based on samples drawn from each. This is a two-sample-problem, and our goal is to test whether $\mathbb{P} = \mathbb{Q}$, using the MMD, an integral probability metric. The MMD uses directly the Kernel Mean Embedding introduced in section 1.2 in order to compare the two distributions.
Formally, we set up the problem as:
For two probability measures \(\mathbb{P}\) and \(\mathbb{Q}\) defined on a metric space \(\mathcal{X}\), \(\mathbb{P} = \mathbb{Q} \iff \mathbb{E}_\mathbb{P}[f(x)] = \mathbb{E}_\mathbb{Q}[f(x)] \; \forall f \in C(\mathcal{X})\), \(C(\mathcal{X})\) being the set of continuous bounded functions on \(\mathcal{X}\). We now define the test-statistic based on the class of functions in the unit ball of a Reproducing Kernel Hilbert Space \(\mathcal{H}_k\) as:
Definition (Maximum Mean Discrepancy).
\[\text{MMD}_k(\mathbb{P},\mathbb{Q} ) := \sup_{\substack{f \in \mathcal{H}_k \\ \|f\|_{\mathcal{H}_k} \leq 1}} \left( \mathbb{E}_{X \sim \mathbb{P}}[f(x)] - \mathbb{E}_{Y \sim \mathbb{Q}}[f(y)] \right)\]The restriction to the unit ball \(\|f\|_{\mathcal{H}_k} \leq 1\) serves two purposes. It prevents the supremum from being infinite by bounding the function class; and more importantly, it ensures that the MMD measures genuine distributional differences rather than differences that could be amplified by arbitrary scaling. The unit ball constraint makes the MMD a proper metric, satisfying the triangle inequality to provide a meaningful notion of distance.
Using the reproducing property, and writing \(\text{MMD}_k(\mathbb{P},\mathbb{Q} )\) as \(\text{MMD}_k\), we obtain:
\[\text{MMD}_k = \sup_{\substack{f \in \mathcal{H}_k \\ \|f\|_{\mathcal{H}_k} \leq 1}} \langle f, \mu_\mathbb{P} - \mu_\mathbb{Q} \rangle_{\mathcal{H}_k} = \|\mu_\mathbb{P} - \mu_\mathbb{Q}\|_{\mathcal{H}_k}\]This result follows from the Cauchy-Schwarz inequality, where the supremum of \(\langle f, g \rangle_{\mathcal{H}_k}\) over unit vectors $f$ is achieved when $f$ is aligned with $g$. So the MMD can be simply defined as the RKHS norm of the difference between mean embeddings. If the distributions are identical, their mean embeddings should coincide and the MMD should be 0.
Expanding the squared MMD, we can now obtain the kernel-based expression:
\[\begin{align*} \text{MMD}^2_k &= \|\mu_\mathbb{P} - \mu_\mathbb{Q}\|^2_{\mathcal{H}_k} \\ &= \langle \mu_\mathbb{P} - \mu_\mathbb{Q}, \mu_\mathbb{P} - \mu_\mathbb{Q} \rangle_{\mathcal{H}_k} \\ &= \langle \mu_\mathbb{P}, \mu_\mathbb{P} \rangle_{\mathcal{H}_k} - 2\langle \mu_\mathbb{P}, \mu_\mathbb{Q} \rangle_{\mathcal{H}_k} + \langle \mu_\mathbb{Q}, \mu_\mathbb{Q} \rangle_{\mathcal{H}_k} \\ &= \mathbb{E}_{X,X' \sim \mathbb{P}}[k(X, X')] - 2\mathbb{E}_{X \sim \mathbb{P}, Y \sim \mathbb{Q}}[k(X, Y)] + \mathbb{E}_{Y,Y' \sim \mathbb{Q}}[k(Y, Y')] \end{align*}\]In practice, we work with the squared MMD for computational purposes. We can estimate it directly from samples without having to compute the feature maps, or the mean embeddings explicitly. This is another instance of the kernel trick, where we can use kernel evaluations instead of manually computing the inner products between potentially infinite-dimensional feature maps.
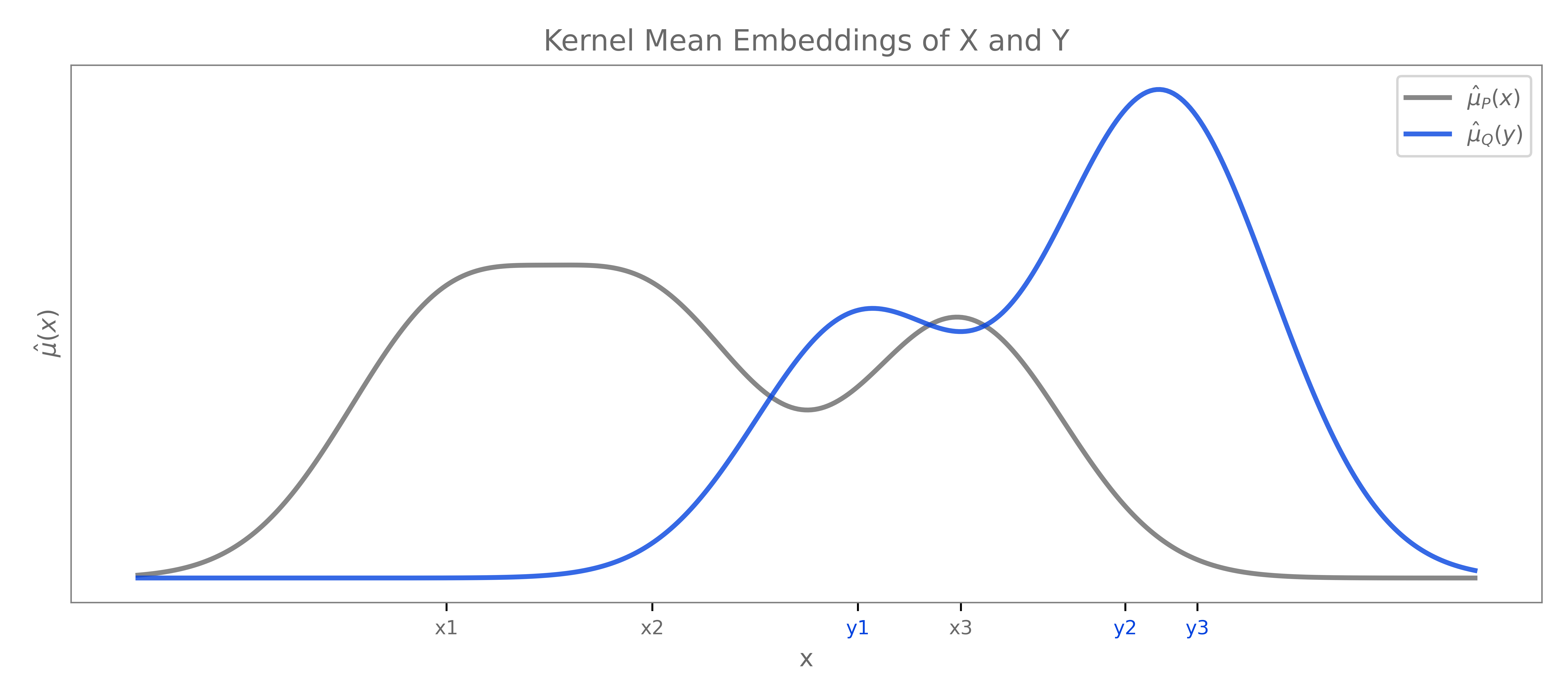
Theorem (Characteristic Kernels and MMD). If $k$ is a characteristic kernel on \(\mathcal{X}\), then \(\text{MMD}_k(\mathbb{P}, \mathbb{Q}) = 0\) if and only if \(\mathbb{P} = \mathbb{Q}\). Thus, for characteristic kernels, the MMD defines a proper metric on the space of probability measures.
This theorem essentially guarantees the validity of the test-statistic. We can reject the null hypothesis \(\mathbb{P} = \mathbb{Q}\) based on a large MMD value.
2.1.2 Empirical Estimation
Given finite samples \(\{x_1, \ldots, x_m\} \sim \mathbb{P}\) and \(\{y_1, \ldots, y_n\} \sim \mathbb{Q}\), we need to estimate \(\text{MMD}^2_k(\mathbb{P}, \mathbb{Q})\) from data. Substituting the empirical mean embeddings into the quadratic form expression yields the natural empirical estimator.
Biased V-Statistic Estimator
The V-statistic is obtained by directly plugging the empirical mean embeddings into the population MMD$^2$ formula:
\[\widehat{\text{MMD}}_v^2 = \frac{1}{m^2} \sum_{i,j=1}^m k(x_i,x_j) - \frac{2}{mn} \sum_{i=1}^{m} \sum_{j=1}^{n} k(x_i,y_j) + \frac{1}{n^2} \sum_{i,j=1}^n k(y_i,y_j).\]This estimator is the most natural empirical analog of the population quantity and has a simple interpretation: it computes the average kernel similarity within each sample and between samples. The V-statistic can be efficiently computed using matrix operations: if \(\mathbf{K}_{XX} \in \mathbb{R}^{m \times m}\), \(\mathbf{K}_{YY} \in \mathbb{R}^{n \times n}\), and \(\mathbf{K}_{XY} \in \mathbb{R}^{m \times n}\) are the kernel Gram matrices, then
\[\widehat{\text{MMD}}_v^2 = \frac{1}{m^2} \mathbf{1}^\top \mathbf{K}_{XX} \mathbf{1} - \frac{2}{mn} \mathbf{1}^\top \mathbf{K}_{XY} \mathbf{1} + \frac{1}{n^2} \mathbf{1}^\top \mathbf{K}_{YY} \mathbf{1}\]Unbiased U-Statistic Estimator
To obtain an unbiased estimator, we exclude the diagonal terms by restricting summations to distinct indices:
\[\widehat{\text{MMD}}_u^2 = \frac{1}{m(m-1)} \sum_{\substack{i,j=1 \\ i \neq j}}^m k(x_i, x_j) - \frac{2}{mn} \sum_{i=1}^{m} \sum_{j=1}^{n} k(x_i, y_j) + \frac{1}{n(n-1)} \sum_{\substack{i,j=1 \\ i \neq j}}^n k(y_i, y_j).\]This estimator is unbiased: \(\mathbb{E}[\widehat{\text{MMD}}_u^2] = \text{MMD}^2(\mathbb{P}, \mathbb{Q})\) exactly for all sample sizes. It is preferred when exact unbiasedness is important, such as in small-sample settings or when theoretical guarantees on Type I error rates are needed. Like the biased estimator, it can be computed efficiently with matrix operations:
\[\widehat{\text{MMD}}_u^2 = \frac{1}{m(m-1)} \mathbf{1}^\top \bar{\mathbf{K}}_{XX} \mathbf{1} - \frac{2}{mn} \mathbf{1}^\top \bar{\mathbf{K}}_{XY} \mathbf{1} + \frac{1}{n(n-1)} \mathbf{1}^\top \bar{\mathbf{K}}_{YY} \mathbf{1},\]where \(\bar{\mathbf{K}}_{XX}\), \(\bar{\mathbf{K}}_{XY}\) and \(\bar{\mathbf{K}}_{YY}\) denote the Gram matrices \(\mathbf{K}_{XX}\), \(\mathbf{K}_{XY}\), and \(\mathbf{K}_{YY}\), with diagonal entries set to 0.
The test thresholds used are found using the Pearson curve (fitting on its first four moments) and bootstrap methods to estimate the distribution of the statistic under the null hypothesis. Gretton et al. (2012) conclude that the Pearson curve method generally provides a more accurate estimate of the threshold for low sample sizes while the two methods provide similar estimates for larger sample sizes, thus making the Bootstrap method preferable as its computational cost is $O(m^2)$ compared to $O(m^3)$ for the Pearson Curves.
2.2 Hilbert-Schmidt Independence Criterion (HSIC)
2.2.1 Theory
The Hilbert-Schmidt Independence Criterion (HSIC) is a kernel-based measure of statistical dependence between two random variables. While MMD tests whether two distributions are identical, HSIC tests whether two random variables are independent by measuring the Hilbert-Schmidt norm of their cross-covariance operator introduced in Section 1.3.2.
For random variables \(X \in \mathcal{X}\) and \(Y \in \mathcal{Y}\) with joint distribution \(\mathbb{P}_{XY}\) and marginals \(\mathbb{P}_X\) and \(\mathbb{P}_Y\), independence is characterized by \(\mathbb{P}_{XY} = \mathbb{P}_X \otimes \mathbb{P}_Y\). The cross-covariance operator $\Sigma_{YX}$ captures the dependence structure between $X$ and $Y$ in the tensor product space \(\mathcal{H}_\mathcal{Y} \otimes \mathcal{H}_\mathcal{X}\).
Definition (Hilbert-Schmidt Independence Criterion). Let \(k_\mathcal{X}\) and \(k_\mathcal{Y}\) be kernels on \(\mathcal{X}\) and \(\mathcal{Y}\) with RKHSs \(\mathcal{H}_\mathcal{X}\) and \(\mathcal{H}_\mathcal{Y}\). The HSIC between $X$ and $Y$ is defined as the squared Hilbert-Schmidt norm of the associated cross-covariance operator $\Sigma_{YX}$:
\[\text{HSIC}(X, Y) := \|\Sigma_{YX}\|_{\text{HS}}^2\]where \(\Sigma_{YX}: \mathcal{H}_\mathcal{X} \to \mathcal{H}_\mathcal{Y}\) is the cross-covariance operator.
Remark (Hilbert-Schmidt Norm and Singular Values). For a compact Hilbert-Schmidt operator \(C: \mathcal{H} \to \mathcal{G}\) with singular value decomposition \(C = \sum_{i=1}^\infty \sigma_i \langle \cdot, h_i \rangle_{\mathcal{H}} g_i\), where \(\{\sigma_i\}_{i=1}^\infty\) are the singular values and \((h_i)_{i \in I}\), \((g_i)_{i \in I}\) are orthonormal bases of \(\mathcal{H}\) and \(\mathcal{G}\) respectively, the Hilbert-Schmidt norm can equivalently be expressed as:
\[\|C\|_{\text{HS}}^2 = \sum_{i \in I} \|Ch_i\|_{\mathcal{G}}^2 = \sum_{i=1}^\infty \sigma_i^2(C)\]The Hilbert-Schmidt norm aggregates information across all modes of the operator, capturing the total magnitude of the linear transformation. In the context of HSIC, the Hilbert-Schmidt norm measures the magnitude of the cross-covariance operator. When $X$ and $Y$ are independent, \(\Sigma_{YX} = 0\) and thus \(\text{HSIC}(X,Y) = 0\).
Using \(\Sigma_{YX} = \mathbb{E}_{YX}[\phi(x) \otimes \psi(y)] - \mu_{\mathbb{P}_X} \otimes \mu_{\mathbb{P}_Y}\), we can expand the Hilbert-Schmidt norm using the definition of the cross-covariance operator, and obtain the kernel-based expression:
\[\begin{align*} \text{HSIC}(X,Y) = \|\Sigma_{YX}\|_{\text{HS}}^2 &= \langle \:\mathbb{E}_{YX}[\phi(x) \otimes \psi(y)] - \mu_{\mathbb{P}_X} \otimes \mu_{\mathbb{P}_Y} \:, \: \mathbb{E}_{YX}[\phi(x) \otimes \psi(y)] - \mu_{\mathbb{P}_X} \otimes \mu_{\mathbb{P}_Y} \:\rangle \\ &= \mathbb{E}_{XY, X'Y'}\big[k_\mathcal{X}(X,X')k_\mathcal{Y}(Y,Y')\big] + \mathbb{E}_{X,X'}\big[k_\mathcal{X}(X,X')\big] \mathbb{E}_{Y,Y'}\big[k_\mathcal{Y}(Y,Y')\big] \\ &\qquad - 2\mathbb{E}_{XY}\big[\mathbb{E}_{X'}\big[k_\mathcal{X}(X,X')\big] \mathbb{E}_{Y'}\big[k_\mathcal{Y}(Y,Y')\big]\big] \end{align*}\]where the expectations are taken over independent copies $(X,Y)$ and $(X’,Y’)$. This expression involves only kernel evaluations and expectations, enabling empirical estimation without explicit feature map computations. This is another instance of the kernel trick, allowing us to work directly with kernel evaluations in potentially infinite-dimensional feature spaces.
2.2.2 Empirical Estimation
Given finite paired samples \(\{(x_i, y_i)\}_{i=1}^m\) drawn i.i.d. from \(\mathbb{P}_{XY}\), we estimate \(\text{HSIC}(X,Y)\) from data. Following the MMD framework, we can derive both biased and unbiased estimators based on the empirical cross-covariance operator.
Biased V-Statistic Estimator
The biased estimator is obtained by directly substituting the empirical cross-covariance operator into the Hilbert-Schmidt norm:
\[\widehat{\text{HSIC}}_v = \frac{1}{m^2} \text{tr}(\mathbf{K}_{XX} \mathbf{H} \mathbf{K}_{YY} \mathbf{H})\]where \(\mathbf{K}_{XX} \in \mathbb{R}^{m \times m}\) and \(\mathbf{K}_{YY} \in \mathbb{R}^{m \times m}\) are the kernel Gram matrices, and \(\mathbf{H} = \mathbf{I} - \frac{1}{m}\mathbf{1}\mathbf{1}^\top\) is the centering matrix.
Similarly to the Maximum Mean Discrepancy estimators, we can also define an unbiased U-statistic estimator of the Hilbert-Schmidt Independence Criterion.
Unbiased U-Statistic Estimator
An unbiased estimator of the HSIC is given by:
\[\widehat{\text{HSIC}}_u = \frac{1}{m(m-3)} \left[ \text{tr}(\bar{\mathbf{K}}_{XX} \bar{\mathbf{K}}_{YY}) + \frac{\mathbf{1}^\top \bar{\mathbf{K}}_{XX} \mathbf{1} \cdot \mathbf{1}^\top \bar{\mathbf{K}}_{YY} \mathbf{1}}{(m-1)(m-2)} - \frac{2}{m-2} \mathbf{1}^\top \bar{\mathbf{K}}_{XX} \bar{\mathbf{K}}_{YY} \mathbf{1} \right]\]where \(\bar{\mathbf{K}}_{XX}\) and \(\bar{\mathbf{K}}_{YY}\) denote the Gram matrices \(\mathbf{K}_{XX}\) and \(\mathbf{K}_{YY}\), respectively, with diagonal entries set to 0. This estimator satisfies \(\mathbb{E}[\widehat{\text{HSIC}}_u] = \text{HSIC}(X,Y)\). The U-statistic is preferred in small-sample settings or when exact unbiasedness is required for hypothesis testing.
2.2.3 Extensions of HSIC
We can use a similar framework to test conditional independence, putting together the HSIC for cross-covariance and the normalized cross-covariance operators introduced in Definition 12. We can then define both the normalized HSIC and the normalized conditional HSIC, following the work and notation of K. Fukumizu, to which we refer the reader for more details, justifications and convergence proofs.
Definition (Conditional HSIC). The conditional HSIC and normalized HSIC are defined as:
\[\begin{align*} I^{COND}(X,Y|Z) &:= \|V_{YX|Z}\|_{\text{HS}}^2\\ I^{NOCCO}(X,Y) &:= \|V_{YX}\|_{\text{HS}}^2 \end{align*}\]where \(V_{YX\|Z} = V_{YX} - V_{YZ}V_{XZ}\) is the normalized conditional cross-covariance operator, and $V_{YX}$ the normalized cross-covariance operator, as defined in section 1.3.2.
For characteristic kernels, \(I^{COND}(X, Y \mid Z) = 0\) if and only if \(X \perp Y \mid Z\), and \(I^{NOCCO}(X,Y) = 0\) if and only if $X \perp Y$. These measures provide a complete characterization of conditional and unconditional independence, respectively.
Empirical Estimation
Given i.i.d. samples \(\{(x_i, y_i, z_i)\}_{i=1}^m\) from the joint distribution \(\mathbb{P}_{XYZ}\), we can construct empirical estimators of both measures. We first define the centered Gram matrices \(\mathbf{\tilde{K}}_{XX}, \mathbf{\tilde{K}}_{YY}, \mathbf{\tilde{K}}_{ZZ} \in \mathbb{R}^{m \times m}\) with entries
\[K_{XX,ij} = \langle k_\mathcal{X}(\cdot, x_i) - \hat{\mu}_{\mathbb{P}_X} \:, \: k_\mathcal{X}(\cdot, x_j) - \hat{\mu}_{\mathbb{P}_X}\rangle_{\mathcal{H}_\mathcal{X}}\]and similarly for \(\mathbf{\tilde{K}}_{YY}\) and \(\mathbf{\tilde{K}}_{ZZ}\), where \(\hat{\mu}_{\mathbb{P}_X}\) are the empirical mean embeddings. To ensure numerical stability, we introduce regularized matrices through the regularization parameter $\epsilon_m > 0$:
\[\mathbf{R}_X = \mathbf{\tilde{K}}_{XX}(\mathbf{\tilde{K}}_{XX} + m\epsilon_m \mathbf{I})^{-1}, \quad \mathbf{R}_{Y} = \mathbf{\tilde{K}}_{YY}(\mathbf{\tilde{K}}_{YY} + m\epsilon_m \mathbf{I})^{-1}, \quad \mathbf{R}_{Z} = \mathbf{\tilde{K}}_{ZZ}(\mathbf{\tilde{K}}_{ZZ} + m\epsilon_m \mathbf{I})^{-1}\]The empirical normalized HSIC is then computed as:
\[\widehat{I}^{NOCCO}_m(X,Y) = \text{tr}(\mathbf{R}_Y \mathbf{R}_X)\]For conditional HSIC, using the empirical normalized conditional cross-covariance operator, the estimator is:
\[\widehat{I}^{COND}_m(X,Y|Z) = \text{tr}(\mathbf{R}_Y \mathbf{R}_X - 2\mathbf{R}_Y \mathbf{R}_X \mathbf{R}_Z + \mathbf{R}_Y \mathbf{R}_Z \mathbf{R}_X \mathbf{R}_Z)\]2.3 Kernel Stein Discrepancy (KSD)
Stein’s method was introduced by James-Stein in the 1970s, but the subsequent developments and applications of his method to non-linear feature maps and more modern statistical learning and hypothesis testing have resurfaced in the 2010s. For this section, we refer mostly to Chwialkowski et al. (2016), Gorham & Mackey (2015), and Liu et al. (2016) for additional details and proofs on the methods introduced.
2.3.1 Stein’s Method
Stein’s method provides an elegant solution for testing whether samples are drawn from a target distribution, in the case where the target is only known up to a normalizing constant, which might be intractable to compute. Classical goodness-of-fit tests, such as Kolmogorov-Smirnov or likelihood-based methods required access to the normalized target density. Maximum Mean Discrepancy requires samples from both distributions, which we may not have for the target.
Stein’s method relies on an identity that eliminates the dependence on the normalizing constant. We first introduce the Stein Operator.
Definition (Stein Operator). For a smooth probability density $p$ on \(\mathbb{R}^d\), the Stein operator $\mathcal{T}_p$ is the linear operator acting on smooth vector-valued test functions \(f : \mathbb{R}^d \to \mathbb{R}^d\) defined by:
\[\mathcal{T}_p f(x) := \nabla_x \log p(x) \cdot f(x) + \nabla_x \cdot f(x),\]where \(\nabla_x \log p(x) \cdot f(x) = \sum_{i=1}^d \frac{\partial \log p}{\partial x_i} f_i(x)\) denotes the Euclidean inner product, and $\nabla_x \cdot f(x) = \sum_{i=1}^d \frac{\partial f_i}{\partial x_i}$ is the divergence of $f$.
The crucial property of this operator is that the score function $\nabla_x \log p(x)$ depends only on the unnormalized density. This operator is particularly useful because it depends on $p$ only through the score function \(\nabla_x \log p(x)\). If \(p(x) = \tilde{p}(x)/Z\) where $Z$ is an intractable normalizing constant, then:
\[\nabla_x \log p(x) = \nabla_x \log \frac{\tilde{p}(x)}{Z} = \nabla_x \log \tilde{p}(x) - \nabla_x \log Z = \nabla_x \log \tilde{p}(x)\]since $Z$ is a constant. Thus, the identity can be evaluated without knowledge of the normalizing constant. We now define Stein’s identity:
Theorem (Stein’s Identity). Let $p$ be a smooth probability density on \(\mathbb{R}^d\) with \(\nabla_x p(x) \to 0\) as \(\|x\| \to \infty\). For any smooth vector-valued function \(f : \mathbb{R}^d \to \mathbb{R}^d\) that decays sufficiently fast at infinity,
\[\mathbb{E}_{x \sim p}\left[ \mathcal{T}_p f(x) \right] = 0\]Proof sketch (1d case).
\[\begin{align*} \mathbb{E}_{x \sim p}\left[\frac{p'(x)}{p(x)} f(x) + f'(x)\right] &= \int_{-\infty}^{\infty} \left[p'(x) f(x) + p(x) f'(x)\right] dx \\ &= \int_{-\infty}^{\infty} \frac{d}{dx}[p(x) f(x)] dx \\ &= [p(x) f(x)]_{-\infty}^{\infty} = 0 \end{align*}\]where the last equality follows from the boundary conditions $p(x) \to 0$ as $x \to \pm\infty$. The multivariate case follows by applying this argument component-wise. $\square$
Stein’s identity provides a principled approach to test discrepancies without requiring a normalizing constant nor samples from the target distribution. If samples are drawn from $p$, then \(\mathbb{E}_{x \sim p}[\mathcal{T}_p f(x)] = 0\) for all suitable $f$. Conversely, given samples \(\{x_i\}_{i=1}^n\) from an unknown distribution $q$, we can evaluate \(\mathbb{E}_{x \sim q}[\mathcal{T}_p f(x)]\), which we expect to be $0$ in the case where $q = p$. The magnitude of the Stein discrepancy \(\mathbb{E}_{x \sim q}[\mathcal{T}_p f(x)]\) should provide evidence against the hypothesis that $q = p$. This raises the question of the class of functions one should consider for $f$. We’ll see in the next section that we can apply a procedure similar to the framework of the Maximum Mean Discrepancy statistic in section 2.1.1, using a kernel-based approach by considering all functions in the unit ball of a RKHS.
2.3.2 KSD
Following the framework of the Maximum Mean Discrepancy, we adopt a kernel-based approach by considering all functions in the unit ball of a reproducing kernel Hilbert space.
Let \(\mathcal{H}^d = \mathcal{H} \times \cdots \times \mathcal{H}\) be the product RKHS of vector-valued functions \(f: \mathbb{R}^d \to \mathbb{R}^d\), where \(\mathcal{H}\) is an RKHS with kernel \(k: \mathbb{R}^d \times \mathbb{R}^d \to \mathbb{R}\).
Using the derivative reproducing property (Proposition 3) on the product space $\mathcal{H}^d$, note that:
\[\nabla^\top f(x) = \langle f, \nabla k(x, \cdot)\rangle_{\mathcal{H}^d} \qquad \text{and} \qquad f(x)^\top \nabla \log p(x) = \langle f, \nabla \log p(x) k(x, \cdot )\rangle_{\mathcal{H}^d} \qquad \forall f\in \mathcal{H}^d, x\in \mathbb{R}^d\]We can now express the Stein operator as an inner product within $\mathcal{H}^d$ using once more the reproducing property:
\[\mathcal{T}_p f(x) = \nabla_x \log p(x) \cdot f(x) + \nabla_x \cdot f(x) = \langle f, \nabla \log p(x) k(x, \cdot ) +\nabla k(x, \cdot)\rangle_{\mathcal{H}^d}\]And following the notation of Chwialkowski et al. (2016), writing \(\xi_p (x, \cdot) := \nabla \log p(x) k(x, \cdot ) +\nabla k(x, \cdot)\), we can now define the expected Stein operator for samples from a distribution $q$ as an inner product in the RKHS:
\[\mathbb{E}_{x \sim q}\left[ \mathcal{T}_p f(x) \right] = \mathbb{E}_{x \sim q}\left[\langle f, \xi_p (x, \cdot) \rangle \right] = \langle f, \mathbb{E}_{x \sim q}\left[\xi_p (x, \cdot)\right] \rangle\]We finally define the Kernelized Stein Discrepancy as the supremum of the expected Stein operator over the class of all functions in the unit ball of the RKHS $\mathcal{H}^d$.
Definition (Kernelized Stein Discrepancy). For a distribution $q$ on $\mathbb{R}^d$ and a target distribution $p$, the Kernelized Stein Discrepancy is defined as:
\[\text{KSD}(q, p) := \sup_{\substack{f \in \mathcal{H}^d \\ \|f\|_{\mathcal{H}^d} \leq 1}} \mathbb{E}_{x \sim q}[\mathcal{T}_p f(x)] = \sup_{\substack{f \in \mathcal{H}^d \\ \|f\|_{\mathcal{H}^d} \leq 1}} \langle f, \mathbb{E}_{x \sim q}\left[\xi_p (x, \cdot)\right] \rangle = \| \mathbb{E}_{x \sim q}\left[\xi_p (x, \cdot) \right]\|_{\mathcal{H}^d}\]using Cauchy-Schwarz, similarly as for the derivation of the closed-form of the Maximum Mean Discrepancy in section 2.1.1.
We can now express the squared KSD as:
\[\text{KSD}^2(q, p) = \| \mathbb{E}_{x \sim q}\left[\xi_p (x, \cdot) \right]\|_{\mathcal{H}^d}^2 = \mathbb{E}_{x \sim q, x' \sim q}\left[ h_P(x, x') \right]\]where $h_P$ is the Stein Kernel, defined as:
\[\begin{align*} h_P(x, x') &:= \nabla \log p(x)^\top\nabla \log p(x') k(x, x') \\ &\quad+ \nabla \log p(x')^\top\nabla_x k(x, x') \\ &\quad+ \nabla \log p(x)^\top\nabla_{x'} k(x, x') \\ &\quad+ \langle \nabla_x k(x, \cdot), \nabla_{x'} k(\cdot, x')\rangle_{\mathcal{H}^d} \end{align*}\]Once again, this enables us to use the “kernel trick” in order to evaluate the (squared) Kernel Stein Discrepancy using only kernel evaluations and the score function $\nabla_x \log p(x)$ of the unnormalized target density.
2.3.3 Empirical Estimation
Similarly to the Maximum Mean Discrepancy and Hilbert-Schmidt Independence Criterion chapters, we now focus on the empirical estimation of the Kernel Stein Discrepancy statistic from data samples. Given finite samples ${x_1, \ldots, x_m}$ drawn i.i.d. from an unknown distribution $q$, we can estimate $\text{KSD}^2(q, p)$ from data using empirical averages of the Stein kernel.
Biased V-Statistic Estimator
The V-statistic estimator is obtained by directly substituting the empirical distribution into the population formula:
\[\widehat{\text{KSD}}_v^2 = \frac{1}{m^2}\sum_{i,j=1}^m h_P(x_i, x_j)\]This estimator is the empirical analog of \(\text{KSD}^2(q, p) = \mathbb{E}_{x, x' \sim q}[h_P(x, x')]\). The V-statistic includes diagonal terms $h(x_i, x_i)$ and is therefore biased for finite samples. Setting $\mathbf{H}_{XX}$ as the kernel Gram matrix of the Stein kernel $h_P$, we can also express it in matrix form:
\[\widehat{\text{KSD}}_v^2 = \frac{1}{m^2} \mathbf{1}^\top \mathbf{H}_{XX}\mathbf{1}\]Unbiased U-Statistic Estimator
To obtain an unbiased estimator, we exclude the diagonal terms, denoting by \(\mathbf{\tilde{H}}_{XX}\) the Stein Kernel Gram matrix \(\mathbf{H}_{XX}\) with its diagonal elements set to 0:
\[\widehat{\text{KSD}}_u^2 = \frac{1}{m(m-1)}\sum_{\substack{i,j=1 \\ i\neq j}}^m h_P(x_i, x_j) \qquad \text{and its matrix form} \quad \widehat{\text{KSD}}_u^2 = \frac{1}{m(m-1)} \mathbf{1}^\top \mathbf{\tilde{H}}_{XX}\mathbf{1}\]This estimator satisfies $\mathbb{E}[\widehat{\text{KSD}}_u^2] = \text{KSD}^2(q, p)$. Similarly to the MMD and HSIC U-statistics, this estimator is preferred when exact unbiasedness is required, such as in small-sample settings.
To test the null hypothesis $H_0: q = p$ versus $H_1: q \neq p$, we compare the empirical KSD against a threshold, generally defined as a quantile of its distribution over the null-hypothesis, obtained via the Wild Bootstrap method.
References
Theory of Reproducing Kernels (1950)
Nachman Aronszajn
Paper
Reproducing Kernel Hilbert Spaces in Probability and Statistics (2011)
Alain Berlinet, Christine Thomas-Agnan
Book
A Kernel Test of Goodness of Fit (2016)
Kacper Chwialkowski, Heiko Strathmann, Arthur Gretton
Paper
Dimensionality Reduction for Supervised Learning with Reproducing Kernel Hilbert Spaces (2004)
Kenji Fukumizu, Francis R. Bach, Michael I. Jordan
Paper
Kernel Measures of Conditional Dependence (2007)
Kenji Fukumizu, Arthur Gretton, Xiaohai Sun, Bernhard Schölkopf
Paper
Kernel Dimension Reduction in Regression (2009)
Kenji Fukumizu, Francis R. Bach, Michael I. Jordan
Paper
Measuring Sample Quality with Stein’s Method (2015)
Jackson Gorham, Lester Mackey
Paper
A Kernel Method for the Two-Sample Problem (2007)
Arthur Gretton, Karsten M. Borgwardt, Malte Rasch, Bernhard Schölkopf, Alexander Smola
Paper
Measuring Statistical Dependence with Hilbert-Schmidt Norms (2005)
Arthur Gretton, Olivier Bousquet, Alex Smola, Bernhard Schölkopf
Paper
A Kernel Two-Sample Test (2012)
Arthur Gretton, Karsten M. Borgwardt, Malte J. Rasch, Bernhard Schölkopf, Alexander Smola
Paper
A Kernelized Stein Discrepancy for Goodness-of-fit Tests (2016)
Qiang Liu, Jason Lee, Michael Jordan
Paper
Methods of Modern Mathematical Physics I: Functional Analysis (1980)
Michael Reed, Barry Simon
Book
Learning with Kernels (1998)
Alexander J. Smola, Bernhard Schölkopf
Book
A Hilbert Space Embedding for Distributions (2007)
Alex Smola, Arthur Gretton, Le Song, Bernhard Schölkopf
Paper
Supervised Feature Selection via Dependence Estimation (2007)
Le Song, Alex Smola, Arthur Gretton, Karsten M. Borgwardt, Justin Bedo
Paper
Hilbert Space Embeddings and Metrics on Probability Measures (2010)
Bharath K. Sriperumbudur, Arthur Gretton, Kenji Fukumizu, Bernhard Schölkopf, Gert R. G. Lanckriet
Paper
A Bound for the Error in the Normal Approximation to the Distribution of a Sum of Dependent Random Variables (1972)
Charles Stein
Paper
Support Vector Machines (2008)
Ingo Steinwart, Andreas Christmann
Book
Derivative Reproducing Properties for Kernel Methods in Learning Theory (2008)
Ding-Xuan Zhou
Paper